How Many Million Of Years Do Scientists Believe Separate Animal And Plant Globin Genes
In this and the preceding 3 chapters, we discussed the structure of genes, the way they are arranged in chromosomes, the intricate cellular machinery that converts genetic information into functional poly peptide and RNA molecules, and the many ways in which gene expression is regulated by the cell. In this section, we discuss some of the means that genes and genomes have evolved over time to produce the vast diversity of mod-solar day life forms on our planet. Genome sequencing has revolutionized our view of this process of molecular evolution, uncovering an astonishing wealth of information about the family relationships among organisms and evolutionary mechanisms.
Information technology is perhaps not surprising that genes with similar functions tin can be found in a diverse range of living things. Only the great revelation of the past 20 years has been the discovery that the actual nucleotide sequences of many genes are sufficiently well conserved that homologous genes—that is, genes that are similar in their nucleotide sequence because of a mutual ancestry—tin often be recognized across vast phylogenetic distances. For case, unmistakable homologs of many man genes are piece of cake to notice in such organisms as nematode worms, fruit flies, yeasts, and even bacteria.
As discussed in Affiliate iii and again in Chapter 8, the recognition of sequence homology has become a major tool for inferring gene and protein role. Although finding such a homology does not guarantee similarity in function, it has proven to be an excellent clue. Thus, it is often possible to predict the function of a gene in humans for which no biochemical or genetic information is available only by comparing its sequence to that of an intensively studied factor in another organism.
Factor sequences are often far more tightly conserved than is overall genome construction. Every bit discussed in Chapter iv, features of genome system such equally genome size, number of chromosomes, gild of genes along chromosomes, abundance and size of introns, and corporeality of repetitive DNA are found to differ greatly amidst organisms, every bit does the actual number of genes.
The number of genes is merely very roughly correlated with the phenotypic complication of an organism. Thus, for example, current estimates of factor number are vi,000 for the yeast Saccharomyces cerevisiae, 18,000 for the nematode Caenorhabditis elegans, 13,000 for Drosophila melanogaster, and thirty,000 for humans (come across Tabular array 1-1). As nosotros shall soon meet, much of the increment in cistron number with increasing biological complexity involves the expansion of families of closely related genes, an observation that establishes gene duplication and difference every bit major evolutionary processes. Indeed, it is likely that all present-mean solar day genes are descendants—via the processes of duplication, departure, and reassortment of gene segments—of a few ancestral genes that existed in early life forms.
Genome Alterations are Acquired by Failures of the Normal Mechanisms for Copying and Maintaining DNA
With a few exceptions, cells do not have specialized mechanisms for creating changes in the structures of their genomes: evolution depends instead on accidents and mistakes. Most of the genetic changes that occur result simply from failures in the normal mechanisms by which genomes are copied or repaired when damaged, although the motion of transposable DNA elements also plays an important role. Every bit we discussed in Chapter 5, the mechanisms that maintain Deoxyribonucleic acid sequences are remarkably precise—simply they are not perfect. For example, because of the elaborate Dna-replication and Dna-repair mechanisms that enable DNA sequences to exist inherited with extraordinary fidelity, merely about ane nucleotide pair in a thousand is randomly changed every 200,000 years. Nevertheless, in a population of x,000 individuals, every possible nucleotide substitution will have been "tried out" on about 50 occasions in the course of a million years—a short span of time in relation to the evolution of species.
Errors in DNA replication, Dna recombination, or Deoxyribonucleic acid repair tin lead either to unproblematic changes in Deoxyribonucleic acid sequence—such as the substitution of ane base pair for another—or to large-scale genome rearrangements such as deletions, duplications, inversions, and translocations of DNA from ane chromosome to another. It has been argued that the rates of occurrence of these mistakes accept themselves been shaped by evolutionary processes to provide an adequate balance between genome stability and change.
In add-on to failures of the replication and repair machinery, the various mobile Dna elements described in Chapter 5 are an of import source of genomic change. In particular, transposable Dna elements (transposons) play a major office as parasitic Dna sequences that colonize a genome and tin spread inside it. In the process, they ofttimes disrupt the function or alter the regulation of existing genes; and sometimes they fifty-fifty create altogether novel genes through fusions between transposon sequences and segments of existing genes. Examples of the iii major classes of transposons were presented in Table v-3, p. 287. Over long periods of evolutionary time, these transposons have profoundly affected the structure of genomes.
The Genome Sequences of Two Species Differ in Proportion to the Length of Time That They Accept Separately Evolved
The differences betwixt the genomes of species live today accept accumulated over more than 3 billion years. Lacking a directly record of changes over time, nosotros can still reconstruct the process of genome evolution from detailed comparisons of the genomes of gimmicky organisms.
The basic tool of comparative genomics is the phylogenetic tree. A simple instance is the tree describing the divergence of humans from the great apes (Figure vii-108). The primary back up for this tree comes from comparisons of gene and protein sequences. For case, comparisons between the sequences of man genes or proteins and those of the great apes typically reveal the fewest differences betwixt human being and chimpanzee and the nigh betwixt human and orangutan.

Figure 7-108
A phylogenetic tree showing the human relationship between the homo and the groovy apes based on nucleotide sequence information. As indicated, the sequences of the genomes of all iv species are estimated to differ from the sequence of the genome of their terminal common (more...)
For closely related organisms such as humans and chimpanzees, it is possible to reconstruct the factor sequences of the extinct, last mutual ancestor of the ii species (Figure 7-109). The close similarity between human being and chimpanzee genes is mainly due to the short time that has been available for the accumulation of mutations in the two diverging lineages, rather than to functional constraints that have kept the sequences the aforementioned. Evidence for this view comes from the observation that even Dna sequences whose nucleotide order is functionally unconstrained—such every bit the sequences that lawmaking for the fibrinopeptides (run into p. 236) or the third position of "synonymous" codons (codons specifying the aforementioned amino acid—see Figure 7-109)—are nigh identical.
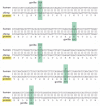
Figure 7-109
Tracing the antecedent sequence from a sequence comparison of the coding regions of human and chimpanzee leptin genes. Leptin is a hormone that regulates food intake and energy utilization in response to the adequacy of fat reserves. As indicated past the (more...)
For less closely related organisms such as humans and mice, the sequence conservation found in genes is largely due to purifying selection (that is, selection that eliminates individuals carrying mutations that interfere with important genetic functions), rather than to an inadequate time for mutations to occur. As a upshot, protein-coding sequences and regulatory sequences in the Deoxyribonucleic acid that are constrained to engage in highly specific interactions with conserved proteins are often remarkably conserved. In contrast, almost DNA sequences in the human being and mouse genomes have diverged so far that it is often incommunicable to marshal them with i another.
Integration of phylogenetic trees based on molecular sequence comparisons with the fossil tape has led to the best available view of the evolution of modern life forms. The fossil record remains important as a source of absolute dates based on the disuse of radioisotopes in the rock formations in which fossils are found. Even so, precise divergence times between species are difficult to establish from the fossil record even for species that leave skilful fossils with distinctive morphology. Populations may be small and geographically localized for long periods before a newly arisen species expands in numbers sufficiently to leave a fossil record that is detectable. Furthermore, fifty-fifty when a fossil closely resembles a gimmicky species, it is not certain that information technology is ancestral to information technology—the fossil may come up from an extinct lineage, while the truthful ancestors of the contemporary species may remain unknown.
The integrated phylogenetic trees support the basic thought that changes in the sequences of detail genes or proteins occur at a constant charge per unit, at least in the lineages of organisms whose generation times and overall biological characteristics are quite similar to one another. This apparent constancy in the rates at which sequences change is referred to as the molecular-clock hypothesis. As described in Affiliate 5, the molecular clock runs most rapidly in sequences that are not subject area to purifying pick—such as intergenic regions, portions of introns that lack splicing or regulatory signals, and genes that have been irreversibly inactivated by mutation (the then-called pseudogenes). The clock runs most slowly for sequences that are bailiwick to strong functional constraints—for case, the amino acid sequences of proteins such every bit actin that appoint in specific interactions with big numbers of other proteins and whose construction, therefore, is highly constrained (meet, for instance, Figure 16-fifteen).
Considering molecular clocks run at rates that are determined both by mutation rates and by the amount of purifying selection on particular sequences, a different calibration is required for genes replicated and repaired by dissimilar systems inside cells. Most notably, clocks based on functionally unconstrained mitochondrial Deoxyribonucleic acid sequences run much faster than clocks based on functionally unconstrained nuclear sequences because of the high mutation rate in mitochondria.
Molecular clocks have a finer time resolution than the fossil record and are a more reliable guide to the detailed structure of phylogenetic trees than are classical methods of tree structure, which are based on comparisons of the morphology and development of dissimilar species. For case, the precise relationship among the cracking-ape and human lineages was non settled until sufficient molecular-sequence data accumulated in the 1980s to produce the tree that was shown in Figure 7-108.
The Chromosomes of Humans and Chimpanzees Are Very Similar
We have just seen that the extent of sequence similarity between homologous genes in different species depends on the length of time that has elapsed since the two species last had a common ancestor. The same principle applies to the larger scale changes in genome structure.
The human and chimpanzee genomes—with their v-million-year history of separate development—are still nigh identical in overall arrangement. Non only practise humans and chimpanzees announced to have essentially the same gear up of xxx,000 genes, but these genes are arranged in nearly the same way along the chromosomes of the two species (encounter Effigy 4-57). The only substantial exception is that man chromosome 2 arose by a fusion of two chromosomes that are dissever in the chimpanzee, the gorilla, and the orangutan.
Even the massive resculpting of genomes that can exist produced past transposon activity has had only minor furnishings on the five-million-yr time scale of the human-chimpanzee departure. For example, more than than 99% of the ane one thousand thousand copies of the Alu family unit of retrotransposons that are present in both genomes are in corresponding positions. This observation indicates that almost of the Alu sequences in our genome underwent duplication and transposition before the difference of the homo and chimpanzee lineages. Nevertheless, the Alu family is even so actively transposing. Thus, a small number of cases accept been observed in which new Alu insertions have caused human genetic affliction; these cases involve transposition of this Deoxyribonucleic acid into sites unoccupied in the genomes of the patient'south parents. More than mostly, at that place exists a class of "man-specific" Alu sequences that occupy sites in the human genome that are unoccupied in the chimpanzee genome. Since perfect-excision mechanisms for Alu sequences appear to be lacking, these man-specific Alu sequences most likely reflect new insertions in the homo lineage, rather than deletions in the chimpanzee lineage. The close sequence similarity among all of the man-specific Alu sequences suggests that they have a contempo mutual ancestor; it may even be that only a single "primary" Alu sequence remains capable of spawning new copies of itself in humans.
A Comparing of Human and Mouse Chromosomes Shows How The Large-scale Structures of Genomes Diverge
The human and chimpanzee genomes are much more akin than are the human and mouse genomes. Although the size of the mouse genome is approximately the same and it contains about identical sets of genes, at that place has been a much longer time period over which changes have had a gamble to accumulate—approximately 100 one thousand thousand years versus 5 million years. It may likewise be that rodents accept significantly college mutation rates than humans; in this case the not bad divergence of the human and mouse genomes would be dominated by a high rate of sequence change in the rodent lineage. Lineage-specific differences in mutation rates are, however, difficult to estimate reliably, and their contribution to the patterns of sequence difference observed among gimmicky organisms remains controversial.
As indicated by the DNA sequence comparison in Figure 7-110, mutation has led to extensive sequence divergence between humans and mice at all sites that are not nether selection—such as the nucleotide sequences of introns. Indeed, human-mouse-sequence comparisons are much more informative of the functional constraints on genes than are human-chimpanzee comparisons. In the latter case, about all sequence positions are the same simply considering non enough time has elapsed since the last common ancestor for big numbers of changes to have occurred. In contrast, considering of functional constraints in human-mouse comparisons the exons in genes stand out as small islands of conservation in a bounding main of introns.

Effigy vii-110
Comparison of a portion of the mouse and homo leptin genes. Positions where the sequences differ by a single nucleotide substitution are boxed in green, and positions that differ past the addition or deletion of nucleotides are boxed in yellow. Note that (more than...)
As the number of sequenced genomes increases, comparative genome analysis is becoming an increasingly important method for identifying their functionally important sites. For example, conservation of open-reading frames between distantly related organisms provides much stronger testify that these sequences are really the exons of expressed genes than does a computational analysis of any one genome. In the future, detailed biological notation of the sequences of complex genomes—such as those of the human and the mouse—will depend heavily on the identification of sequence features that are conserved beyond multiple, distantly related mammalian genomes.
In contrast to the state of affairs for humans and chimpanzees, local cistron order and overall chromosome organization have diverged greatly between humans and mice. According to rough estimates, a total of nigh 180 break-and-rejoin events have occurred in the homo and mouse lineages since these two species last shared a common ancestor. In the process, although the number of chromosomes is similar in the two species (23 per haploid genome in the homo versus 20 in the mouse), their overall structures differ greatly. For instance, while the centromeres occupy relatively primal positions on most human chromosomes, they lie next to an cease of each chromosome in the mouse. Nevertheless, fifty-fifty subsequently the all-encompassing genomic shuffling, there are many big blocks of Dna in which the cistron order is the same in the human and the mouse. These regions of conserved factor order in chromosomes are referred to as synteny blocks (run into Effigy iv-18).
Analysis of the transposon families in the human and the mouse provide additional evidence of the long divergence time separating the two species. Although the major retrotransposon families in the human have counterparts in the mouse—for example, human Alu repeats are similar in sequence and transposition mechanism to the mouse B1 family unit—the two families have undergone split up expansions in the two lineages. Even in regions where human and mouse sequences are sufficiently conserved to allow reliable alignment, there is no correlation betwixt the positions of Alu elements in the man genome and the B1 elements in corresponding segments of the mouse genome (Effigy seven-111).

Figure 7-111
A comparison of the β-globin cistron cluster in the human and mouse genomes, showing the location of transposable elements. This stretch of man genome contains five functional β-globin-like genes (orange); the comparable region from the (more...)
It Is Difficult to Reconstruct the Structure of Ancient Genomes
The genomes of ancestral organisms can be inferred, but never directly observed: there are no ancient organisms alive today. Although a modern organism such equally the horseshoe crab looks remarkably similar to fossil ancestors that lived 200 million years ago, there is every reason to believe that the horseshoe-crab genome has been changing during all that time at a rate similar to that occurring in other evolutionary lineages. Selective constraints must accept maintained key functional backdrop of the horseshoe-crab genome to business relationship for the morphological stability of the lineage. However, genome sequences reveal that the fraction of the genome subject to purifying selection is pocket-size; hence the genome of the modern horseshoe crab must differ greatly from that of its extinct ancestors, known to us simply through the fossil tape.
Information technology is difficult to infer even gross features of the genomes of long-extinct organisms. An important example is the so-chosen introns-early versus introns-late controversy. Soon after the discovery in 1977 that the coding regions of well-nigh genes in metazoan organisms are interrupted by introns, a argue arose about whether introns reflect a late conquering during the evolution of life on earth or whether they were instead present in the primeval genes. According to the introns-early model, fast-growing organisms such as leaner lost the introns present in their ancestors because they were under option for a compact genome adapted for rapid replication. This view is contested past an introns-late model, in which introns are viewed as having been inserted into intronless genes long afterwards the evolution of unmarried-jail cell organisms, mayhap through the bureau of certain types of transposons.
At that place is presently no reliable way of resolving this controversy. Comparative studies of existing genomes provide estimates of rates of intron proceeds and loss in various evolutionary lineages. All the same, these estimates bear only indirectly on the question of how genomes were organized billions of years agone. Bacteria and humans are equally "modern" organisms, both of whose genomes differ and then profoundly from that of their last mutual antecedent that we tin can only speculate about the properties of this very aboriginal, ancestral genome.
When two modern organisms share about identical patterns of intron positions in their genes, we can be confident that the introns were present in the last common ancestor of the two species. An illuminating comparison involves humans and the puffer fish, Fugu rubripes (Effigy 7-112). The Fugu genome is remarkable in having an unusually small size for a vertebrate (0.4 billion nucleotide pairs compared to one billion or more for many other fish and 3 billion for typical mammals). The small size of the Fugu genome is due almost entirely to the modest size of its introns. Specifically, Fugu introns, likewise every bit other non-coding segments of the Fugu genome, lack the repetitive Deoxyribonucleic acid that makes up a large portion of the genomes of nearly well studied vertebrates. However, the positions of Fugu introns are almost perfectly conserved relative to their positions in mammalian genomes (Figure 7-113).
Figure seven-112
The puffer fish, Fugu rubripes. (Courtesy of Byrappa Venkatesh.)

Figure seven-113
Comparison of the genomic sequences of the human and Fugu genes encoding the poly peptide huntingtin. Both genes (indicated in cherry) contain 67 short exons that align in one:1 correspondence to one another; these exons are connected by curved lines. The human being (more...)
The question of why Fugu introns are so small is reminiscent of the introns-early versus introns-belatedly argue. Plainly, either introns grew in many lineages while staying small-scale in the Fugu lineage, or the Fugu lineage experienced massive loss of repetitive sequences from its introns. Nosotros have a clear agreement of how genomes tin grow past agile transposition since most transposition events are duplicative [i.due east., the original re-create stays where it was while a copy inserts at the new site (run into Figures 5-72 and v-76)]. There is considerably less evidence in well-studied organisms for mutational processes that would efficiently delete transposons from immense numbers of sites without likewise deleting side by side functionally critical sequences at rates that would threaten the survival of the lineage. Nonetheless, the origin of Fugu's unusually small introns remains uncertain.
Cistron Duplication and Divergence Provide a Critical Source of Genetic Novelty During Evolution
Much of our give-and-take of genome evolution then far has emphasized neutral change processes or the effects of purifying selection. However, the near important characteristic of genome development is the capacity for genomic modify to create biological novelty that tin can be positively selected for during evolution, giving rise to new types of organisms.
Comparisons between organisms that seem very different illuminate some of the sources of genetic novelty. A hit characteristic of these comparisons is the relative scarcity of lineage-specific genes (for case, genes found in primates but not in rodents, or those found in mammals just not in other vertebrates). Much more prominent are selective expansions of preexisting factor families. The genes encoding nuclear hormone receptors in humans, a nematode worm, and a fruit fly, all of which have fully sequenced genomes, illustrate this signal (Effigy 7-114). Many of the subtypes of these nuclear receptors (besides called intracellular receptors) have shut homologs in all three organisms that are more like to each other than they are to other family subtypes present in the same species. Therefore, much of the functional divergence of this large cistron family unit must accept preceded the deviation of these iii evolutionary lineages. Subsequently, one major branch of the cistron family underwent an enormous expansion simply in the worm lineage. Similar, simply smaller lineage-specific expansions of particular subtypes are evident throughout the cistron family tree, but they are specially evident in the homo—suggesting that such expansions offer a path toward increased biological complexity.

Figure vii-114
A phylogenetic tree based on the inferred protein sequences for all nuclear hormone receptors encoded in the genomes of homo (H. sapiens), a nematode worm (C. elegans), and a fruit fly (D. melanogaster). Triangles represent protein subfamilies that (more...)
Gene duplication appears to occur at high rates in all evolutionary lineages. An exam of the affluence and rate of departure of duplicated genes in many different eucaryotic genomes suggests that the probability that any detail gene volition undergo a successful duplication event (i.east., one that spreads to near or all individuals in a species) is approximately i% every one thousand thousand years. Piffling is known near the precise mechanism of gene duplication. However, because the ii copies of the gene are ofttimes adjacent to one another immediately following duplication, it is thought that the duplication frequently results from inexact repair of double-strand chromosome breaks (see Effigy v-53).
Duplicated Genes Diverge
A major question in genome evolution concerns the fate of newly duplicated genes. In most cases, at that place is presumed to be footling or no selection—at least initially—to maintain the duplicated state since either copy tin provide an equivalent function. Hence, many duplication events are probable to be followed by loss-of-function mutations in 1 or the other cistron. This cycle would functionally restore the one-cistron state that preceded the duplication. Indeed, there are many examples in contemporary genomes where one copy of a duplicated gene can be seen to have become irreversibly inactivated past multiple mutations. Over time, the sequence similarity between such a pseudogene and the functional gene whose duplication produced it would be expected to be eroded past the aggregating of many mutational changes in the pseudogene—eventually becoming undetectable.
An alternative fate for gene duplications is for both copies to remain functional, while diverging in their sequence and design of expression and taking on dissimilar roles. This procedure of "duplication and difference" nearly certainly explains the presence of large families of genes with related functions in biologically circuitous organisms, and information technology is thought to play a critical role in the evolution of increased biological complexity.
Whole-genome duplications offer peculiarly dramatic examples of the duplication-divergence bicycle. A whole-genome duplication can occur quite only: all that is required is 1 circular of genome replication in a germline cell lineage without a corresponding cell division. Initially, the chromosome number simply doubles. Such abrupt increases in the ploidy of an organism are mutual, peculiarly in fungi and plants. After a whole-genome duplication, all genes exist equally duplicate copies. Nonetheless, unless the duplication upshot occurred so recently that there has been picayune time for subsequent alterations in genome structure, the results of a series of segmental duplications—occurring at unlike times—are very hard to distinguish from the terminate production of a whole-genome duplication. In the instance of mammals, for example, the office of whole genome duplications versus a serial of piecemeal duplications of DNA segments is quite uncertain. Nevertheless, it is articulate that a bang-up deal of factor duplication has ocurred in the distant past.
Analysis of the genome of the zebrafish, in which either a whole-genome duplication or a series of more than local duplications occurred hundreds of millions of years agone, has cast some light on the process of gene duplication and divergence. Although many duplicates of zebrafish genes appear to take been lost past mutation, a pregnant fraction—perhaps as many as xxx–50%—have diverged functionally while both copies have remained active. In many cases, the most obvious functional difference betwixt the duplicated genes is that they are expressed in different tissues or at different stages of evolution (run into Figure 21-45). Ane attractive theory to explain such an end result imagines that unlike, mildly deleterious mutations apace occur in both copies of a duplicated factor set up. For example, 1 copy might lose expression in a particular tissue due to a regulatory mutation, while the other re-create loses expression in a second tissue. Following such an occurrence, both gene copies would be required to provide the full range of functions that were once supplied by a single gene; hence, both copies would now exist protected from loss through inactivating mutations. Over a longer flow of time, each copy could then undergo further changes through which it could acquire new, specialized features.
The Development of the Globin Gene Family Shows How Dna Duplications Contribute to the Development of Organisms
The globin cistron family unit provides a peculiarly adept case of how DNA duplication generates new proteins, because its evolutionary history has been worked out particularly well. The unmistakable homologies in amino acid sequence and structure amongst the nowadays-solar day globins indicate that they all must derive from a common bequeathed gene, even though some are now encoded by widely separated genes in the mammalian genome.
We can reconstruct some of the past events that produced the diverse types of oxygen-carrying hemoglobin molecules by because the different forms of the protein in organisms at different positions on the phylogenetic tree of life. A molecule similar hemoglobin was necessary to permit multicellular animals to grow to a large size, since large animals could no longer rely on the elementary diffusion of oxygen through the torso surface to oxygenate their tissues fairly. Consequently, hemoglobin-like molecules are found in all vertebrates and in many invertebrates. The most primitive oxygen-carrying molecule in animals is a globin polypeptide chain of about 150 amino acids, which is found in many marine worms, insects, and primitive fish. The hemoglobin molecule in college vertebrates, however, is composed of two kinds of globin chains. It appears that about 500 million years ago, during the evolution of college fish, a series of gene mutations and duplications occurred. These events established 2 slightly different globin genes, coding for the α- and β-globin chains in the genome of each individual. In modernistic higher vertebrates each hemoglobin molecule is a complex of 2 α bondage and 2 β chains (Figure 7-115). The four oxygen-binding sites in the α2β2 molecule collaborate, assuasive a cooperative allosteric change in the molecule as it binds and releases oxygen, which enables hemoglobin to take up and to release oxygen more than efficiently than the single-chain version.

Figure seven-115
A comparison of the structure of one-chain and four-chain globins. The four-concatenation globin shown is hemoglobin, which is a complex of 2 α- and β-globin bondage. The one-concatenation globin in some archaic vertebrates forms a dimer that dissociates (more...)
Notwithstanding later, during the evolution of mammals, the β-chain gene apparently underwent duplication and mutation to give ascension to a second β-like chain that is synthesized specifically in the fetus. The resulting hemoglobin molecule has a higher affinity for oxygen than adult hemoglobin and thus helps in the transfer of oxygen from the female parent to the fetus. The gene for the new β-similar chain after mutated and duplicated again to produce 2 new genes, ε and γ, the ε chain being produced before in development (to form αiiε2) than the fetal γ concatenation, which forms α2γ2. A duplication of the adult β-chain gene occurred notwithstanding after, during primate evolution, to give rise to a δ-globin factor and thus to a minor form of hemoglobin (αiiδ2) constitute only in developed primates (Figure 7-116).
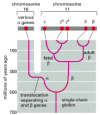
Figure vii-116
An evolutionary scheme for the globin chains that carry oxygen in the blood of animals. The scheme emphasizes the β-like globin cistron family. A relatively contempo gene duplication of the γ-chain gene produced γG and γA, which (more...)
Each of these duplicated genes has been modified by point mutations that affect the properties of the last hemoglobin molecule, too every bit by changes in regulatory regions that determine the timing and level of expression of the cistron. As a effect, each globin is made in unlike amounts at dissimilar times of homo development (see Figure 7-60B).
The end result of the gene duplication processes that accept given rising to the diversity of globin chains is seen clearly in the human genes that arose from the original β gene, which are bundled equally a serial of homologous DNA sequences located within 50,000 nucleotide pairs of 1 some other. A similar cluster of α-globin genes is located on a divide man chromosome. Because the α- and β-globin gene clusters are on dissever chromosomes in birds and mammals but are together in the frog Xenopus, it is believed that a chromosome translocation upshot separated the two gene clusters about 300 1000000 years ago (meet Figure 7-116).
There are several duplicated globin DNA sequences in the α- and β-globin gene clusters that are not functional genes, but pseudogenes. These have a close homology to the functional genes but have been disabled past mutations that foreclose their expression. The existence of such pseudogenes make it clear that, as expected, not every DNA duplication leads to a new functional factor. We likewise know that nonfunctional Deoxyribonucleic acid sequences are not rapidly discarded, equally indicated past the large excess of noncoding Deoxyribonucleic acid that is establish in mammalian genomes.
Genes Encoding New Proteins Can Be Created by the Recombination of Exons
The role of Dna duplication in evolution is not confined to the expansion of factor families. It tin can also act on a smaller calibration to create single genes by stringing together short, duplicated segments of DNA. The proteins encoded by genes generated in this mode tin can be recognized past the presence of repeating, like protein domains, which are covalently linked to one another in series. The immunoglobulins (Figure 7-117) and albumins, for example, too as most fibrous proteins (such equally collagens) are encoded by genes that have evolved by repeated duplications of a primordial DNA sequence.
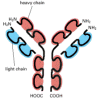
Figure seven-117
Schematic view of an antibiotic (immunoglobulin) molecule. This molecule is a complex of two identical heavy bondage and ii identical light chains. Each heavy chain contains four like, covalently linked domains, while each low-cal concatenation contains two such (more...)
In genes that have evolved in this style, as well every bit in many other genes, each separate exon frequently encodes an individual protein folding unit, or domain. It is believed that the system of Dna coding sequences every bit a serial of such exons separated by long introns has greatly facilitated the evolution of new proteins. The duplications necessary to form a unmarried gene coding for a protein with repeating domains, for case, can occur by breaking and rejoining the DNA anywhere in the long introns on either side of an exon encoding a useful protein domain; without introns there would be only a few sites in the original gene at which a recombinational exchange between Deoxyribonucleic acid molecules could indistinguishable the domain. Past enabling the duplication to occur by recombination at many potential sites rather than just a few, introns increase the probability of a favorable duplication upshot.
More generally, we know from genome sequences that component parts of genes—both their individual exons and their regulatory elements—have served equally modular elements that have been duplicated and moved most the genome to create the present great diversity of living things. As a result, many present-day proteins are formed as a patchwork of domains from dissimilar domain families, reflecting their long evolutionary history (Figure 7-118).

Figure 7-118
Domain structure of a grouping of evolutionary related proteins that are thought to have a similar office. In general, at that place is a tendency for the proteins in more complex organisms, such every bit ourselves, to contain additional domains—as is the example (more...)
Genome Sequences Have Left Scientists with Many Mysteries to Exist Solved
Now that we know from genome sequences that a homo and a mouse contain essentially the same genes, we are forced to confront ane of the major bug that will challenge jail cell biologists throughout the adjacent century. Given that a human and a mouse are formed from the same gear up of proteins, what has happened during the evolutionary process to make a mouse and a human so different? Although the answer is present somewhere among the three billion nucleotides in each sequenced genome, we do not yet know how to decipher this blazon of information—then that the answer to this critical, most fundamental question is not known.
Despite our ignorance, it is perhaps worth engaging in a bit of speculation, if only to assistance signal the way forward to some of the hard issues ahead. In biology, timing is everything, as volition become clear when we examine the elaborate mechanisms that allow a fertilized egg to develop into an embryo, and the embryo to develop into an adult (discussed in Chapter 21). The human body is formed equally the result of many billions of decisions that are made during our development equally to which RNA molecule and which protein are to exist made where, every bit well every bit exactly when and in what amount each is to be produced. These decisions are unlike for a human than for a chimpanzee or a mouse. The coding sequences of genomes represent a more or less standard prepare of the 30,000 or so basic parts from which all 3 organisms are fabricated. It is therefore the many different types of controls on factor expression described in this Chapter that must largely create the difference between a human and other mammals.
Given these assumptions, information technology would exist reasonable to wait genomes to accept evolved in a way that allows organisms to experiment with contradistinct gene timing and expression patterns in selected cells. We have already seen some show that this is so, when we discussed alternative RNA splicing and RNA editing mechanisms. There besides appear to be mechanisms—some based on the movements of transposable DNA elements—that let modules to exist readily added to and subtracted from the regulatory regions of genes, so as to produce changes in the pattern of their transcription as organisms evolve. In fact, an analysis of these regulatory regions provides bear witness to support the claim that most gene regulatory regions have been formed by the evolutionary mixing and matching of the Deoxyribonucleic acid-binding sites that are recognized past gene regulatory proteins (Figure 7-119).
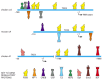
Figure vii-119
Cistron command regions for mouse and chicken eye lens crystallins. Crystallins make up the bulk of the lens and are responsible for refracting and focusing calorie-free onto the retina. Many proteins in the cell take properties (loftier solubility, proper refractive (more...)
Genetic Variation within a Species Provides a Fine-Scale View of Genome Evolution
In comparisons between two species that have diverged from one some other by millions of years, information technology makes lilliputian difference which individuals from each species are compared. For example, typical human being and chimpanzee Dna sequences differ from i another by 1%. In dissimilarity, when the same region of the genome is sampled from two unlike humans, the differences are typically less than 0.1%. For more distantly related organisms, the inter-species differences overshadow intra-species variation even more dramatically. Nonetheless, each "stock-still deviation" between the man and the chimpanzee (i.e., each difference that is now characteristic of all or nearly all individuals of each species) started out every bit a new mutation in a single private. If the size of the interbreeding population in which the mutation occurred is N, the initial allele frequency of a new mutation would be ½N for a diploid organism. How does such a rare mutation become stock-still in the population, and hence go a characteristic of the species rather than of a particular individual genome?
The answer to this question depends on the functional consequences of the mutation. If the mutation has a significantly deleterious effect, information technology will just be eliminated by purifying selection and will not become fixed. (In the near farthermost case, the individual carrying the mutation will die without producing progeny.) Conversely, the rare mutations that confer a major reproductive reward on individuals who inherit them volition spread rapidly in the population. Considering humans reproduce sexually and genetic recombination occurs each time a gamete is formed, the genome of each individual who has inherited the mutation will exist a unique recombinational mosaic of segments inherited from a large number of ancestors. The selected mutation along with a small corporeality of neighboring sequence—ultimately inherited from the individual in which the mutation occurred—will merely be one piece of this huge mosaic.
The great majority of mutations that are non harmful are non beneficial either. These selectively neutral mutations can also spread and become fixed in a population, and they make a large contribution to the evolutionary modify in genomes. Their spread is not as rapid as the spread of the rare strongly advantageous mutations. The procedure by which such neutral genetic variation is passed down through an idealized interbreeding population tin exist described mathematically by equations that are surprisingly simple. The arcadian model that has proven well-nigh useful for analyzing human genetic variation assumes a abiding population size, and random mating, as well as selective neutrality for the mutations. While neither of these assumptions is a skillful description of man population history, they nonetheless provide a useful starting point for analyzing intra-species variation.
When a new neutral mutation occurs in a abiding population of size North that is undergoing random mating, the probability that it will ultimately become fixed is approximately ½Due north. For those mutations that do become fixed, the average time to fixation is approximately 4N generations. A detailed analysis of data on human genetic variation suggests an ancestral population size of approximately 10,000 during the flow when the current blueprint of genetic variation was largely established. Under these weather condition, the probability that a new, selectively neutral mutation would go fixed was small-scale (5 × ten–v), while the boilerplate time to fixation was on the order of 800,000 years. Thus, while we know that the human population has grown enormously since the development of agriculture approximately fifteen,000 years agone, virtually man genetic variation arose and became established in the man population much earlier than this, when the human being population was still small.
Fifty-fifty though most of the variation among modern humans originates from variation present in a comparatively tiny group of ancestors, the number of variations encountered is very large. Most of the variations take the form of unmarried-nucleotide polymorphisms (SNPs). These are simply points in the genome sequence where one large fraction of the human population has i nucleotide, while another big fraction has some other. Two human genomes sampled from the modern earth population at random volition differ at approximately 2.5 × ten6 sites (1 per 1300 nucleotide pairs). Mapped sites in the human being genome that are polymorphic—pregnant that there is a reasonable probability that the genomes of two individuals volition differ at that site—are extremely useful for genetic analyses, in which ane attempts to associate specific traits (phenotypes) with specific DNA sequences for medical or scientific purposes (run into p. 531).
Confronting the groundwork of ordinary SNPs inherited from our prehistoric ancestors, certain sequences with uncommonly high mutation rates stand out. A dramatic example is provided by CA repeats, which are ubiquitous in the human genome and in the genomes of other eucaryotes. Sequences with the motif (CA)n are replicated with relatively low fidelity because of a slippage that occurs betwixt the template and the newly synthesized strands during Dna replication; hence, the precise value of due north can vary over a considerable range from i genome to the adjacent. These repeats make ideal Dna-based genetic markers, since nigh humans are heterozygous—carrying two values of n at any particular CA repeat, having inherited one repeat length (due north) from their female parent and a dissimilar echo length from their father. While the value of north changes sufficiently rarely that most parent-child transmissions propagate CA repeats faithfully, the changes are sufficiently frequent to maintain high levels of heterozygosity in the human population. These and other uncomplicated repeats that brandish exceptionally high variability provide the footing for identifying individuals by DNA analysis in crime investigations, paternity suits, and other forensic applications (see Figure 8-41).
While most of the SNPs and other mutual variations in the human genome sequence are thought to have no effect on phenotype, a subset of them must be responsible for nearly all of the heritable aspects of human individuality. A major claiming in human genetics is to learn to recognize those relatively few variations that are functionally important—against the big groundwork of neutral variation that distinguishes the genomes of any two human beings.
Summary
Comparisons of the nucleotide sequences of present-day genomes have revolutionized our understanding of gene and genome evolution. Due to the extremely loftier fidelity of DNA replication and DNA repair processes, random errors in maintaining the nucleotide sequences in genomes occur then rarely that merely about 5 nucleotides in yard are contradistinct every million years. Not surprisingly, therefore, a comparison of human and chimpanzee chromosomes—which are separated by about 5 meg years of evolution—reveals very few changes. Non only are our genes essentially the same, but their order on each chromosome is almost identical. In improver, the positions of the transposable elements that brand up a major portion of our noncoding DNA are mostly unchanged.
When one compares the genomes of two more than distantly related organisms—such as a homo and a mouse, separated past most 100 million years—one finds many more than changes. Now the effects of natural selection can be clearly seen: through purifying selection, essential nucleotide sequences—both in regulatory regions and coding sequences (exon sequences)—have been highly conserved. In dissimilarity, nonessential sequences (for example, intron sequences) accept been altered to such an extent that an accurate alignment according to beginnings is often not possible.
Because of purifying selection, homologous genes can be recognized over large phylogenetic distances, and information technology is often possible to construct a detailed evolutionary history of a item cistron, tracing its history dorsum to common ancestors of nowadays-day species. We tin can thereby encounter that a great bargain of the genetic complexity of nowadays-day organisms is due to the expansion of ancient factor families. DNA duplication followed past sequence difference has thus been a major source of genetic novelty during evolution.
Source: https://www.ncbi.nlm.nih.gov/books/NBK26836/
Posted by: breesehicasonfut.blogspot.com

0 Response to "How Many Million Of Years Do Scientists Believe Separate Animal And Plant Globin Genes"
Post a Comment